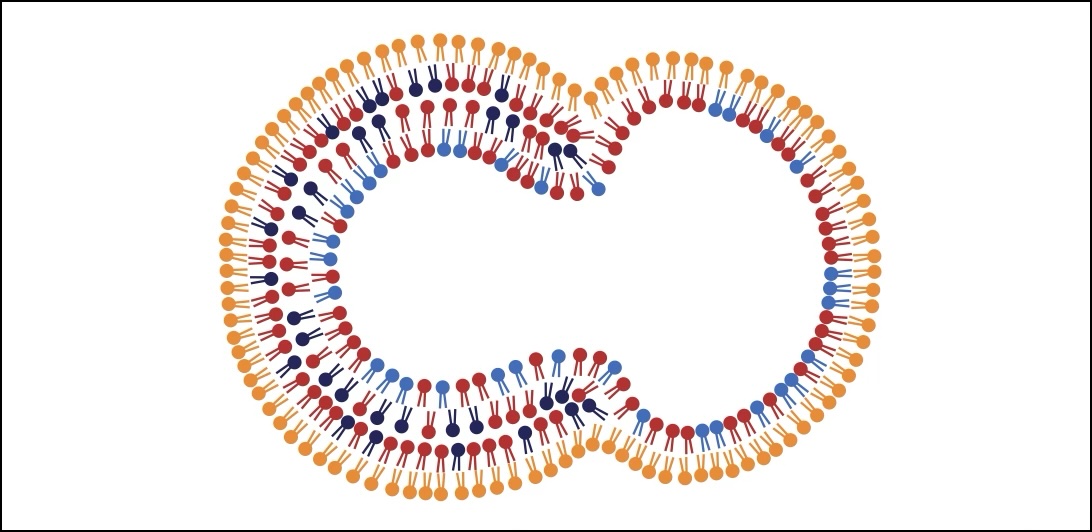
Membrane reshaping is often driven by cytoskeletal filament networks. In collaboration with the Koenderink lab (TU Delft), I investigated the shapes of lipid vesicles containing actin filaments (Baldauf* et al., 2023) and the initiation of actin-driven membrane protrusions (Baldauf et al., 2023). I also contributed to a systematic review with the de Maat lab (University Medical Center Rotterdam) by developing filament network simulations to benchmark image analysis software (de Vries et al., 2023).
References
2023
-
Branched actin cortices reconstituted in vesicles sense membrane curvature
Lucia Baldauf* , Felix Frey* , Marcos Arribas Perez , Timon Idema , and 1 more author
Biophys. J., Feb 2023
*contributed equally
The actin cortex is a complex cytoskeletal machinery that drives and responds to changes in cell shape. It must generate or adapt to plasma membrane curvature to facilitate diverse functions such as cell division, migration, and phagocytosis. Due to the complex molecular makeup of the actin cortex, it remains unclear whether actin networks are inherently able to sense and generate membrane curvature, or whether they rely on their diverse binding partners to accomplish this. Here, we show that curvature sensing is an inherent capability of branched actin networks nucleated by Arp2/3 and VCA. We develop a robust method to encapsulate actin inside giant unilamellar vesicles (GUVs) and assemble an actin cortex at the inner surface of the GUV membrane. We show that actin forms a uniform and thin cortical layer when present at high concentration and distinct patches associated with negative membrane curvature at low concentration. Serendipitously, we find that the GUV production method also produces dumbbell-shaped GUVs, which we explain using mathematical modeling in terms of membrane hemifusion of nested GUVs. We find that branched actin networks preferentially assemble at the neck of the dumbbells, which possess a micrometer-range convex curvature comparable with the curvature of the actin patches found in spherical GUVs. Minimal branched actin networks can thus sense membrane curvature, which may help mammalian cells to robustly recruit actin to curved membranes to facilitate diverse cellular functions such as cytokinesis and migration.
-
Biomimetic actin cortices shape cell-sized lipid vesicles
Lucia Baldauf , Felix Frey , Marcos Arribas Perez , Miroslav Mladenov , and 3 more authors
preprint: bioRxiv, Jan 2023
Animal cells are shaped by a thin layer of actin filaments underneath the plasma membrane known as the actin cortex. This cortex stiffens the cell surface and thus opposes cellular deformation, yet also actively generates membrane protrusions by exerting polymerization forces. It is unclear how the interplay between these two opposing mechanical functions plays out to shape the cell surface. To answer this question, we reconstitute biomimetic actin cortices nucleated by the Arp2/3 complex inside cell-sized lipid vesicles. We show that thin Arp2/3-nucleated actin cortices strongly deform and rigidify the shapes of giant unilamellar vesicles and impart a shape memory on time scales that exceeds the time of actin turnover. In addition, actin cortices can produce finger-like membrane protrusions, showing that Arp2/3-mediated actin polymerization forces alone are sufficient to initiate protrusions in the absence of actin bundling or membrane curving proteins. Combining mathematical modeling and our experimental results reveals that the concentration of actin nucleating proteins, rather than actin polymerization speed, is crucial for protrusion formation. This is because locally concentrated actin polymerization forces can drive a positive feedback loop between recruitment of actin and its nucleators to drive membrane deformation. Our work paints a picture where the actin cortex can either drive or inhibit deformations depending on the local distribution of nucleators.
-
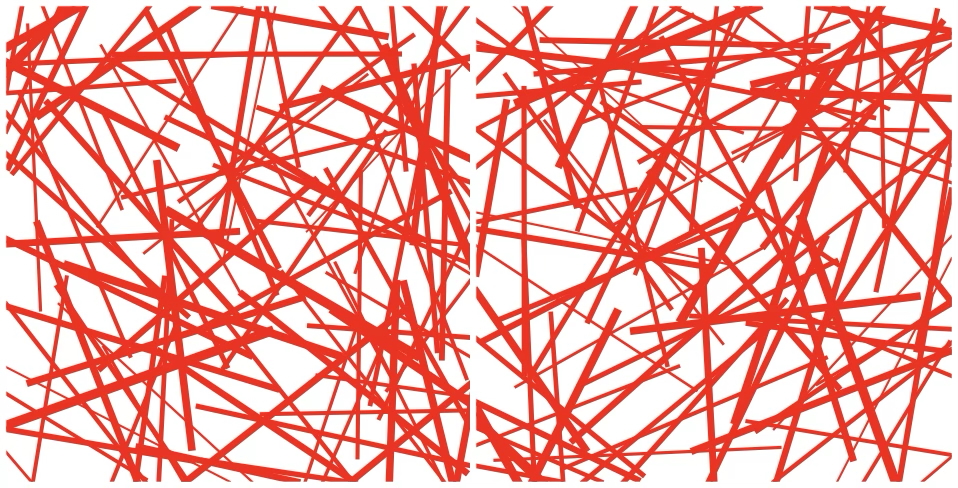
A systematic review and comparison of automated tools for quantification of fibrous networks
Judith J. Vries , Daphne M. Laan , Felix Frey , Gijsje H. Koenderink , and 1 more author
Acta Biomater., Feb 2023
Fibrous networks are essential structural components of biological and engineered materials. Accordingly, many approaches have been developed to quantify their structural properties, which define their material properties. However, a comprehensive overview and comparison of methods is lacking. Therefore, we systematically searched for automated tools quantifying network characteristics in confocal, stimulated emission depletion (STED) or scanning electron microscopy (SEM) images and compared these tools by applying them to fibrin, a prototypical fibrous network in thrombi. Structural properties of fibrin such as fiber diameter and alignment are clinically relevant, since they influence the risk of thrombosis. Based on a systematic comparison of the automated tools with each other, manual measurements, and simulated networks, we provide guidance to choose appropriate tools for fibrous network quantification depending on imaging modality and structural parameter. These tools are often able to reliably measure relative changes in network characteristics, but absolute numbers should be interpreted with care.