Biological cells are enclosed by lipid membranes that must be both sturdy and flexible – to contain the cellular interior and allow membrane reshaping during various cellular processes – such as cell division or endocytosis (the incorporation of cargo into cells). Across nature, different membrane designs have evolved to resolve these partially conflicting requirements. I combined computer simulations and continuum theory to investigate membrane remodeling in archaeal membranes (Amaral* et al., 2025), plant membranes, in collaboration with the Otegui lab (University of Wisconsin-Madison) (Weiner* et al., 2024), and membrane turnover in bio-mimetic systems (Frey & Idema, 2022). I also co-authored a literature review on modeling membrane reshaping (Frey & Idema, 2021) and a tutorial review on mesoscale computer simulations for lipid membranes (Muñoz-Basagoiti* et al., 2025).
References
2025
-
Balancing stability and flexibility when reshaping archaeal membranes
Miguel Amaral* , Felix Frey* , Xiuyun Jiang , Buzz Baum , and 1 more author
eLife, Oct 2025
*contributed equally
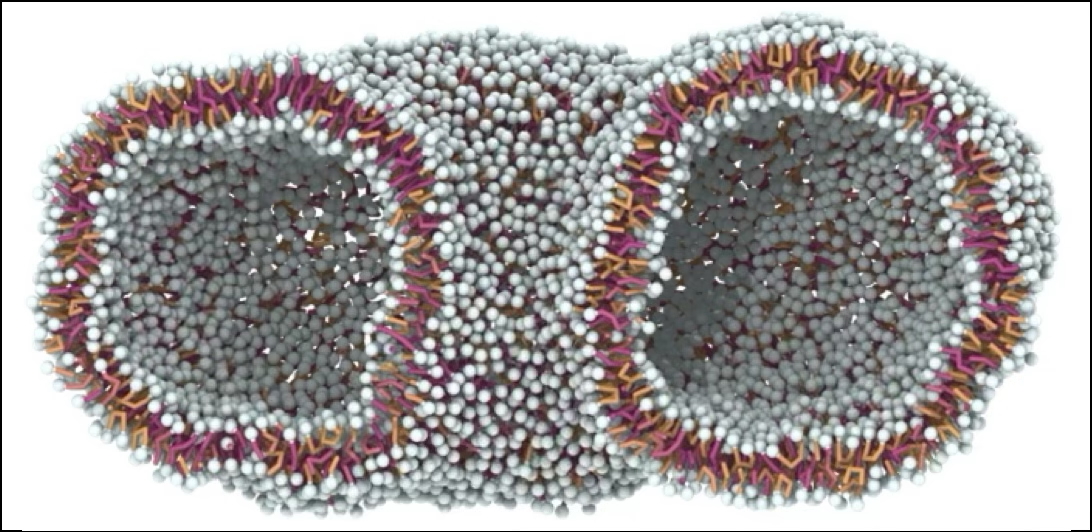
Across the tree of life, distinct designs of cellular membranes have evolved. In bacteria and eukaryotes single-headed lipids self-assemble into flexible bilayer membranes. By contrast, archaea often possess double-headed, monolayer spanning bolalipids, mixed with bilayer lipids, enabling them to survive in harsh environments. Here, using a minimal computational model for bolalipid membranes, we discover trade-offs when forming membranes. We find that membranes made out of flexible bolalipids resemble bilayer membranes as bolalipids exhibit conformational switch into U-shaped conformations to enable higher curvatures. Conversely, stiffer bolalipids, resembling those in extremophile archaea, take on straight conformations and form liquid membranes that are stiff, and prone to pore formation during membrane reshaping. Strikingly, we show how to achieve fluid bolalipid membranes that are both stable and flexible – by including small amounts of bilayer lipids, as archaea do. Our study explains how different organisms resolve trade-offs when generating membranes of desired material properties.
-
A tutorial for mesoscale computer simulations of lipid membranes: tether pulling, tubulation and fluctuations
Maitane Muñoz-Basagoiti* , Felix Frey* , Billie Meadowcroft* , Miguel Amaral* , and 2 more authors
Soft Matter, Jul 2025
*contributed equally
Lipid membranes and membrane deformations are a long-standing area of research in soft matter and biophysics. Computer simulations have complemented analytical and experimental approaches as one of the pillars in the field. However, setting up and using membrane simulations can come with barriers due to the multidisciplinary effort involved and the vast choice of existing simulations models. In this review, we introduce the non-expert reader to coarse-grained membrane simulations (CGMS) at the mesoscale. Firstly, we give a concise overview of the modelling approaches to study fluid membranes, together with guidance to more specialized references. Secondly, we provide a conceptual guide on how to develop CGMS. Lastly, we construct a hands-on tutorial on how to apply CGMS, by providing a pedagogical examination of tether pulling, tubulation and fluctuations with three different membrane models, and discussing them in terms of their scope and how resource-intensive they are. To ease the reader’s venture into the field, we provide a repository with ready-to-run tutorials.
2024
-
Endosomal membrane budding patterns in plants
Ethan Weiner* , Elizabeth Berryman* , Felix Frey* , Ariadna González Solı́s* , and 4 more authors
Proc. Natl. Acad. Sci. U.S.A., Oct 2024
*contributed equally
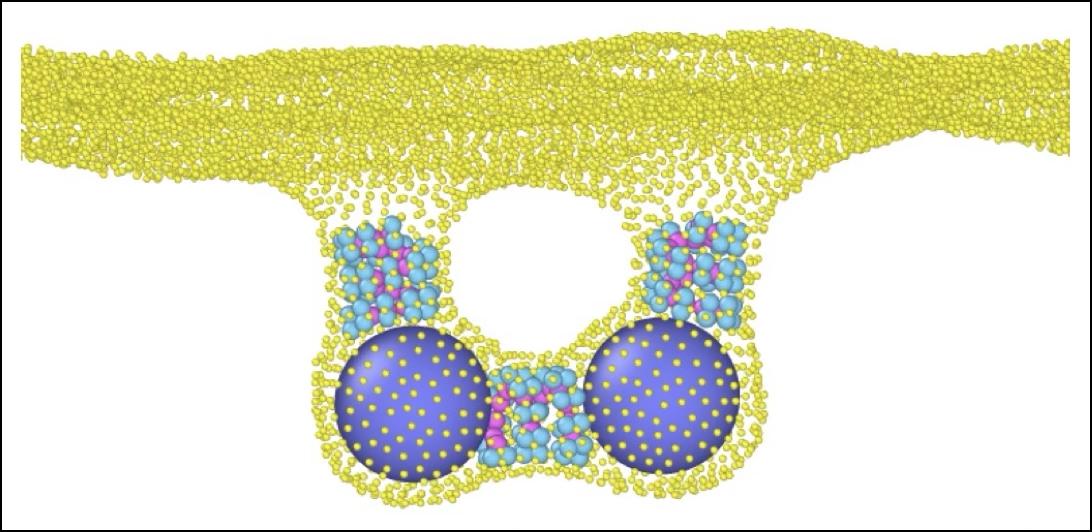
Multivesicular endosomes (MVEs) sequester membrane proteins destined for degradation within intralumenal vesicles (ILVs), a process mediated by the membrane-remodeling action of Endosomal Sorting Complex Required for Transport (ESCRT) proteins. In Arabidopsis, endosomal membrane constriction and scission are uncoupled, resulting in the formation of extensive concatenated ILV networks and enhancing cargo sequestration efficiency. Here, we used a combination of electron tomography, computer simulations, and mathematical modeling to address the questions of when concatenated ILV networks evolved in plants and what drives their formation. Through morphometric analyses of tomographic reconstructions of endosomes across yeast, algae, and various land plants, we have found that ILV concatenation is widespread within plant species, but only prevalent in seed plants, especially in flowering plants. Multiple budding sites that require the formation of pores in the limiting membrane were only identified in hornworts and seed plants, suggesting that this mechanism has evolved independently in both plant lineages. To identify the conditions under which these multiple budding sites can arise, we used particle-based molecular dynamics simulations and found that changes in ESCRT filament properties, such as filament curvature and membrane binding energy, can generate the membrane shapes observed in multiple budding sites. To understand the relationship between membrane budding activity and ILV network topology, we performed computational simulations and identified a set of membrane remodeling parameters that can recapitulate our tomographic datasets.
2022
-
Membrane area gain and loss during cytokinesis
Felix Frey , and Timon Idema
Phys. Rev. E, Aug 2022
In cytokinesis of animal cells, the cell is symmetrically divided into two. Since the cell’s volume is conserved, the projected area has to increase to allow for the change of shape. Here we aim to predict how membrane gain and loss adapt during cytokinesis. We work with a kinetic model in which membrane turnover depends on membrane tension and cell shape. We apply this model to a series of calculated vesicle shapes as a proxy for the shape of dividing cells. We find that the ratio of kinetic turnover parameters changes nonmonotonically with cell shape, determined by the dependence of exocytosis and endocytosis on membrane curvature. Our results imply that controlling membrane turnover will be crucial for the successful division of artificial cells.
2021
-
More than just a barrier: using physical models to couple membrane shape to cell function
Felix Frey , and Timon Idema
Soft Matter, Jan 2021
The correct execution of many cellular processes, such as division and motility, requires the cell to adopt a specific shape. Physically, these shapes are determined by the interplay of the plasma membrane and internal cellular driving factors. While the plasma membrane defines the boundary of the cell, processes inside the cell can result in the generation of forces that deform the membrane. These processes include protein binding, the assembly of protein superstructures, and the growth and contraction of cytoskeletal networks. Due to the complexity of the cell, relating observed membrane deformations back to internal processes is a challenging problem. Here, we review cell shape changes in endocytosis, cell adhesion, cell migration and cell division and discuss how by modeling membrane deformations we can investigate the inner working principles of the cell.